
Goal: molecular microscope, 3D images of individual molecules in situ
Marriage of atomic force microscope (AFM) and MRI imaging
Spins in sample excited by RF exert magnetic force on cantilever, detected by interferometer.
Scan sample past cantilever (or vice versa), deconvolve signal to obtain 3D image.
Invented/developed here at UW School of Medicine and UW Dept. of Mechanical Engineering by John Sidles, Joe Garbini, et al.
RF coil (left), sample positioner (bottom), interferometer (top), cantilever (right)

Cantilever tip with permanent magnet, showing resonant slices due to magnetic field gradient. At a given RF, spins in only one slice exert force on the cantilever.

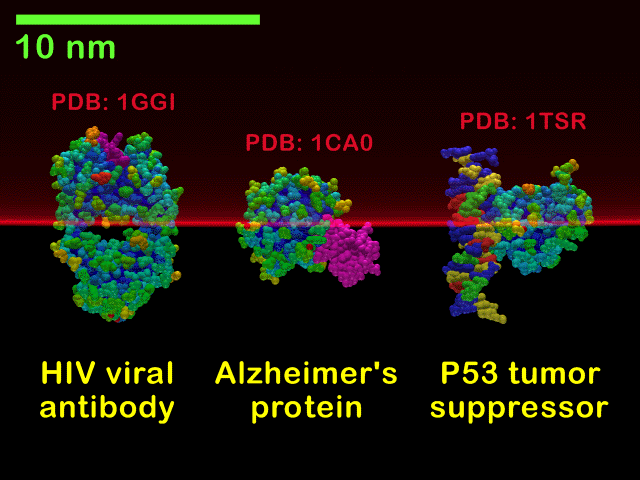
Molecules in a resonant slice
Predict signals and noise expected from molecules of interest: how long does it take for spins to lock to RF, etc?
Investigate coupling between spins in the sample: significant effects?
Investigate MPI-style parallel programming techniques and performance in this application.
| v | spin vector (3) |
| b_tot | magnetic field acting on v (3) |
| b_rf | external RF field (3) |
| B | magnetic coupling between spins (3x3) |
| N | number of spins (10s, 100s, or 1000s) |
| i,j | spin indices |
| dvi/dt | = | b_toti X vi (vector cross product) |
| b_toti | = | b_rfi + Sum (all other j) Bij vj (matrix-vector product) |
Bij describes coupling between spins i and j
Bij is characteristic of the sample (particular molecule),
parameter (input) to simulation
Due to coupling between spins, might be lots of communication --- maybe too much!
Expect computation time proportional to number of interactions, N2
Output is v(t) for all vi, t = 0 1 2 ... tn, tn is 1000s or 10000s
Integrate a system of ODEs
dvi(t)/dt = b_tot X vi(t)
Euler integration: straight-line extrapolation using present rate of change.
vi(t+1) = vi(t) + dt * dvi(t)/dt
Runge-Kutta integration: weighted average of Euler estimates at
different t, dt
We use 4th order Runge-Kutta (4 estimates)
Intuition: each process simulates one spin (or n spins)
N spins, np processes, n = N/np spins/process
Each process just computes, updates one (or n) vi
Due to coupling between spins, each process must read all vj, use all relevant Bij
All updated vi must be distributed to all processes after each time step
In each time step, communication proportional to N, computation proportional to N2
Expect advantage of parallelization to grow with N
BUT there must be an upper limit to the number of significantly interacting spins, independent of N: about 20 or 30 nearest neighbors, we expect.
From scratch, by hand (C and MPI only, no other libraries)
Obvious (naive?) coding: for (i=0;i<n;i++) for (l=0;l<3;l++) v[i][l] = ...
Three stages: read, compute, write. No file I/O during compute stage.
Two test cases: large and small number of spins N = 160, 32. Measure performance of each case on different numbers of processors np = 1,2,4,8,16,32.
Number of spins N and number of time steps tn chosen to take appreciable but not overwhelming time, so timing measurements meaningful. Here is the elapsed time of both cases on a Mac G4 with one processor:
| N | N2 | tn | elapsed | (CPU) | Comment |
| 160 | 25,600 | 1000 | 26 sec | (26) | Large N, parallel advantage expected |
| 32 | 1024 | 25000 | 25 sec | (20) | Typical number of interactions expected |
Not a real molecule, arbitrary (random) initial spins and coupling matrices
Measure multiprocessor performance on ladon cluster. Report elapsed (wall) time and computation (CPU) time (on one CPU) for computation stage only (quite reproducible, unlike read and write) Also report elapsed time only for read and write stages. Elapsed times are in seconds.
With N = 160, the program is pleasingly parallel. Doubling np almost doubles speed up through 8 processors (but little or no improvement after that).
| np | N | tn | read | elapsed | (CPU) | write |
| 1 | 160 | 1000 | 0.28 | 16.66 | (16.62) | 5.23 |
| 2 | 160 | 1000 | 0.40 | 9.41 | ( 8.81) | 5.074 |
| 4 | 160 | 1000 | 0.47 | 6.08 | ( 4.16) | 9.36 |
| 8 | 160 | 1000 | 0.48 | 3.53 | ( 1.84) | 18.26 |
| 16 | 160 | 1000 | 0.72 | 3.38 | ( 1.28) | 7.77 |
| 32 | 160 | 1000 | 1.00 | 3.45 | ( 1.50) | 11.09 |
With N = 32, the program is pathological. Adding processors slows it down!
| np | N | tn | read | elapsed | (CPU) | write |
| 1 | 32 | 25000 | 0.01 | 6.86 | (6.83) | 26.44 |
| 2 | 32 | 25000 | 0.02 | 13.45 | (7.03) | 48.20 |
| 4 | 32 | 25000 | 0.02 | 22.27 | (8.70) | 105.03 |
| 8 | 32 | 25000 | 0.02 | 37.84 | (12.07) | 182.63 |
| 16 | 32 | 25000 | 0.34 | 46.28 | (17.82) | 330.42 |
| 32 | 32 | 25000 | 0.27 | 60.65 | (28.39) | 430.11 |
One processor is enough for small and medium-sized simulations: we can simulate 1000 interactions/msec (~30 spins) on a Mac G4.
The same program can be pleasing or pathological depending on just one parameter; cutover point is hard to predict.
Underwhelming benefit from parallelism in this program, for the two cases tested.
Investigate better parallelization strategy.
Consider 100s or 1000s of spins, where each interacts with ~20 neighbors.
Pre-compute lists of neighbors where interaction matters.
Optimize communications (processes on virtual 3D grid, communicate only with neighbors).
Investigate speeding up serial computation. Vectorization (SSE for x86, Altivec for G4) promises up to 4x speedup (vs 8x found for parallelization).
Visualization (show spins coupled to cantilever)